Tabular
Note
This page describe what is specific to the tabular layout. For more general information creating and using datasets, see Using an existing dataset and Building your own datasets respectively.
Creating
To create a tabular dataset, the layout entry in the recipe’s
output section must be set to tabular:
output:
layout: tabular
Using
To open a tabular dataset, you use the open_dataset function with the start, end, window and frequency parameters.
ds = open_dataset(
dataset,
start=1979,
end=2020,
window="(-6h,0]",
frequency="6h",
)
The default values for start and end are the first and last date
of the dataset, respectively. Because these value may fall on full round
hours, it is recommended to set them explicitly.
Unlike for gridded datasets, the start, end and frequency
parameters can have arbitrary values, and are used to define how windows
are built and how many samples are in the dataset. Note that start
and end can be outside the range of actual dates in the datasets.
When requesting windows outside the range of actual dates, empty records
will be returned to the user.
The default value for window is (-3h,0] and the default value
for frequency is 3h. Windows are relative time intervals
that can be open or closed at each end. A round bracket indicates an
open end, while a square bracket indicates a closed end. The default
units are hours.
Windows can be open or closed at each end:
"[-3,+3]" # Both ends are included
"(-1d,0]" # Start is open, end is closed
Data samples
The dataset is made of samples, which are built by applying the
window to a list of reference dates defined by the start,
end and frequency parameters.
Reference dates
The references dates of the dataset are defined as all dates between
start and end with a step of frequency.
result = []
date = start
while date <= end:
result.append(date)
date += frequency
Note
The reference dates are not necessarily the same as the actual
dates in the dataset. They are used, together with the window
parameter, to define the samples returned when iterating the dataset.
See below for more information. Nevertheless, in order to ensure
compatibility with gridded datasets, the reference dates are
available as the dates attribute of the dataset.
It is not currently possible to combine tabular and gridded datasets within a single call to open_dataset,
but when this will be implemented, ds.dates, ds.frequency,
len(ds), etc. will all be compatible and comparable between the
two layouts.
ds.dates # Returns the list of reference dates defined by start, end and frequency
The number of samples in the dataset is then given by the formula:
number_of_samples = (end - start) // frequency + 1
to get the list of dates, you can access the dates attribute of the
dataset:
ds.dates # Returns the list of reference dates defined by start, end and frequency
The length of the dataset is equal to the number of samples:
assert len(ds) == number_of_samples
Single sample
A sample is a 2D numpy array that is returned when indexing the dataset
with an integer. The first dimension of the array is the number of
observations in the window, and the second dimension is the number of
variables. Each sample is constructed by applying the window to the
corresponding date. For example, if the date is 2020-01-01 00:00:00 and
the window is (-3h,0], then the sample will contain all observations
between 2019-12-31 21:00:00 and 2020-01-01 00:00:00, including the
latter but not the former.
sample = ds[42]
# A 2D array is returned, the first dimension is the number of observations
# in the 43rd window (samples are 0-indexed).
assert len(sample.shape) == 2
# The second dimension is the variables
assert sample.shape[1] == len(ds.variables)
The whole dataset can also be iterated over using a for loop:
for sample in ds:
assert len(sample.shape) == 2
assert sample.shape[1] == len(ds.variables)
is equivalent to:
for i in range(len(ds)):
sample = ds[i]
assert len(sample.shape) == 2
assert sample.shape[1] == len(ds.variables)
Auxiliary information
Becuse tabular data is unstructured, information such as the latidudes, longitudes and dates if the actual data cannot be provided at the dataset level. Instead, it is provided at the sample level. When you access a sample, you can also access the corresponding latitudes, longitudes, dates, etc. This information is returned as attributes of the sample:
Auxiliary information can be accessed as:
sample = ds[42]
number_of_observations_in_window = sample.shape[0]
# Returns the corresponding latitudes
sample.latitudes
assert len(sample.latitudes) == number_of_observations_in_window
# Returns the corresponding longitudes
sample.longitudes
assert len(sample.longitudes) == number_of_observations_in_window
sample.dates # Returns the corresponding row dates
# Returns the corresponding dates
sample.dates
assert len(sample.dates) == number_of_observations_in_window
# Return the reference date of the window
sample.reference_date
assert sample.reference_date == ds.start_date + 42 * ds.frequency
# Return the timedeltas in seconds relative to the reference_date
sample.timedeltas
assert len(sample.timedeltas) == number_of_observations_in_window
Slices
When slicing the dataset, the same rules apply as for indexing with an
integer, but you can recover the samples using the boundaries
attribute of the resulting array . The boudaries attribute is a list
of slice objects that can be used to access the samples in the
result. You also can retrieve the reference dates with the
reference_dates attribute of the result.
samples = ds[10:30]
assert len(samples.boundaries) == 20
assert len(samples.reference_dates) == 20
i = 10
for b in samples.boundaries:
sample = samples[b]
assert np.array_equal(sample, ds[i])
i += 1
The latitudes, longitudes, dates, timedeltas, etc.
attributes of the resulting array are the concatenation of the
corresponding attributes of the samples.
assert np.array_equal(samples.latitudes, np.concatenate([ds[i].latitudes for i in range(10,30)]))
assert np.array_equal(samples.longitudes, np.concatenate([ds[i].longitudes for i in range(10,30)]))
assert np.array_equal(samples.dates, np.concatenate([ds[i].dates for i in range(10,30)]))
Warning
The two codes below are not equivalent:
samples = ds[10:30]
boundaries = samples.boundaries
latitudes = samples.latitudes[boundaries[1]]
and:
samples = ds[10:30]
boundaries = samples.boundaries
latitudes = samples[boundaries[1]].latitudes
Only the first construct will work.
Examples
The following examples show various ways to define the window and the frequency parameters when opening a tabular dataset.
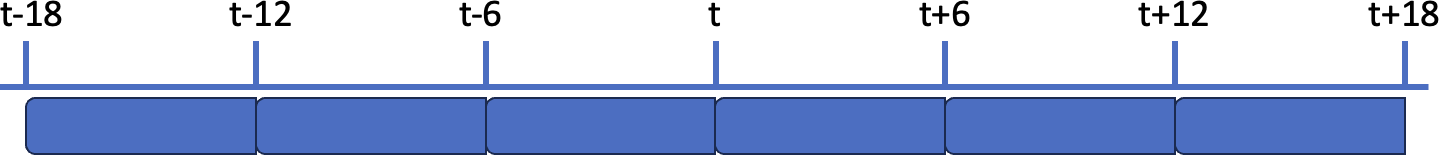
First example, the window width (6h) matches the frequency (6h), so the whole dataset is covered:
ds = open_dataset(
path,
start=1979,
end=2020,
window="(-6h,0]",
frequency="6h")
The schema below illustrates the window and frequency parameters in this case:
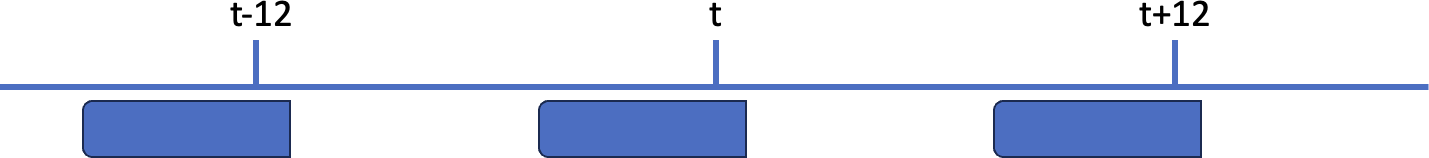
Second example, the window width (6h) is narrower than the frequency (12h), so there are gaps between the windows:
ds = open_dataset(
path,
start=1979,
end=2020,
window="(-5h,+1h]",
frequency="12h")
As illustrated in the schema below, there are gaps between the windows:
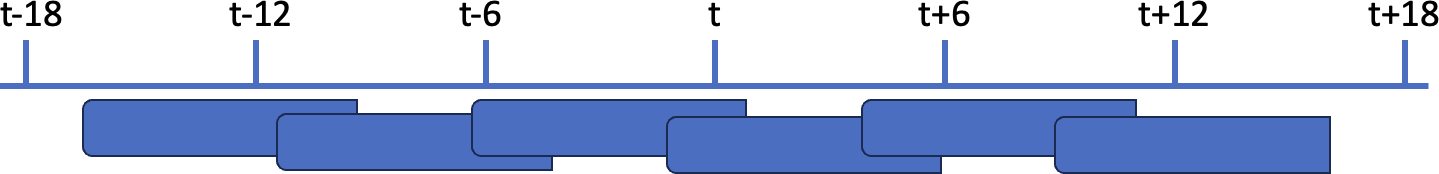
In the third example, the window width (8h) is wider than the frequency (6h), so there are overlaps between the windows:
ds = open_dataset(
path,
start=1979,
end=2020,
window="(-5h,+3h]",
frequency="6h")
As illustrated in the schema below, there are overlaps between the windows: